Network Enhancement
Network Enhancement (NE) is a general method to denoise weighted biological networks. The method improves the signal-to-noise ratio of undirected weighted networks leading to better performance of downstream networks analytics. NE is a principled approach with theoretical guarantees about convergence and performance.
Weighted networks are capable of encoding fine-grained, individual-level, relationships and are thus especially suited for complex networks generated by modern day biological experiments. Due to the expansion of high-throughput biotechnologies, such weighted biological networks are more readily available than ever before.
However, while this expansion in available data presents a great opportunity for network science, the path to knowledge discovery from networks is often obfuscated by experimental and biological noise.
In this project, we develop Network Enhancement (NE), a general method to denoise weighted biological networks. The approach can be incorporated into any weighted network analysis pipeline and can lead to improved downstream analysis.
Network Enhancement as a general method to denoise weighted biological networks.
Bo Wang*, Armin Pourshafeie*, Marinka Zitnik*, Junjie Zhu, Carlos D. Bustamante, Serafim Batzoglou, and Jure Leskovec.
Nature Communications, 9, 3108, 2018. [arXiv]
* Equal contribution.
Network Enhancement method
NE takes as input a noisy, undirected, weighted network and outputs a network on the same set of nodes but with a new set of edge weights.
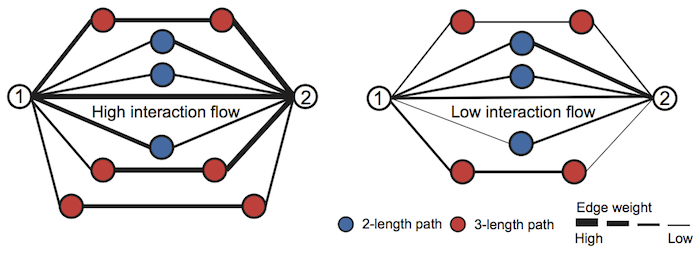
The main crux of NE is the observation that nodes connected through strong (high edge weight) paths in the network are more likely to be linked by a high weight edge. NE employes higher-order network structures to enhance a given weighted biological network. The diffusion process in NE revises edge weights in the network based on interaction flow between any given two nodes, as illustrated in the figure.
Specifically, for any two nodes, NE updates the weight of their edge by modeling all paths of length three or less connecting those nodes.
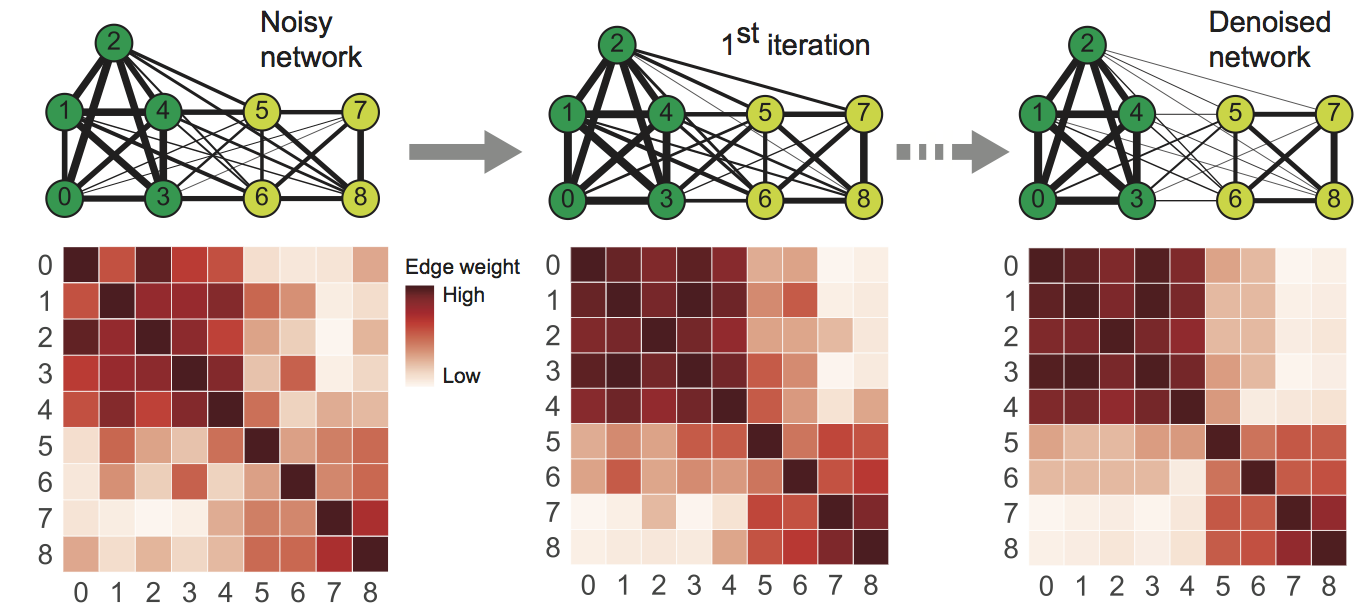
The figure below illustrates the iterative diffusion process of NE. The diffusion process in NE generates a network in which nodes with strong similarity/interactions are connected with high weight edges while nodes with weak similarity/interactions are connected with low weight edges.
Our algorithmic framework for denoising weighted biological networks is:
- Take as input a weighted network and form its associated adjacency matrix (visualized as a heat map below).
- Iteratively update the network using the NE diffusion process.
Upon convergence, the enhanced network has a doubly stochastic matrix property. Mathematically, this means that eigenvectors associated with the input network are preserved while the eigengap is increased. The increased eigengap is a highly appealing property. Among other things, it leads to accurate detection of modules/clusters in network.
Study 1: Human tissue networks
We applied NE to 22 gene interaction networks from different human tissues. The networks capture gene interactions that are specific to human tissues and cell lineages ranging from B lymphocyte to skeletal muscle and the whole brain.
Given an enhanced tissue network, we checked how well relevant tissue-specific gene functions are connected in the network. The expectation is that function-associated genes tend to interact more frequently in tissues in which the function is active than in other non-relevant tissues.
In the NE-enhanced blood plasma network, functions with the highest edge density are blood coagulation, fibrin clot formation, and negative regulation of very-low-density lipoprotein particle remodeling, all these functions are specific to blood plasma (figure below, left). The most connected functions in the NE-enhanced brain network are brain morphogenesis and forebrain regionalization, which are both specific to brain (figure below, right).

Datasets
Raw tissue networks were generated by Greene et al., Nature Genetics 2015. The following table provides links to raw tissue networks in the GIANT database.| File | Description |
|---|---|
| bio-blood-plasma-top.mat | Blood plasma tissue network. |
| bio-blood-platelet-top.mat | Blood platelet tissue network. |
| bio-blood-top.mat | Blood tissue network. |
| bio-blood-vessel-top.mat | Blood vessel tissue network. |
| bio-bone-top.mat | Bone tissue network. |
| bio-brain-top.mat | Brain tissue network. |
| bio-b-lymphocyte-top.mat | B lymphocyte tissue network. |
| bio-cardiac-muscle-top.mat | Cardiac muscle tissue network. |
| bio-central-nervous-system-top.mat | Central nervous system tissue network. |
| bio-chondrocyte-top.mat | Chondrocyte tissue network. |
| bio-embryo-top.mat | Embryo tissue network. |
| bio-epidermis-top.mat | Epidermis tissue network. |
| bio-heart-top.mat | Heart tissue network. |
| bio-lymphocyte-top.mat | Lymphocyte tissue network. |
| bio-muscle-top.mat | Muscle tissue network. |
| bio-natural-killer-cell-top.mat | Natural killer cell tissue network. |
| bio-nervous-system-top.mat | Nervous system tissue network. |
| bio-neuron-top.mat | Neuron tissue network. |
| bio-retina-top.mat | Retina tissue network |
| bio-skeletal-muscle-top.mat | Skeletal muscle tissue network. |
| bio-smooth-muscle-top.mat | Smooth muscle tissue network. |
| bio-t-lymphocyte-top.mat | T lymphocyte tissue network. |
| File | Description |
|---|---|
| bio-blood-plasma-top.mat | Enhanced blood plasma tissue network. |
| bio-blood-platelet-top.mat | Enhanced blood platelet tissue network. |
| bio-blood-top.mat | Enhanced blood tissue network. |
| bio-blood-vessel-top.mat | Enhanced blood vessel tissue network. |
| bio-bone-top.mat | Enhanced bone tissue network. |
| bio-brain-top.mat | Enhanced brain tissue network. |
| bio-b-lymphocyte-top.mat | Enhanced B lymphocyte tissue network. |
| bio-cardiac-muscle-top.mat | Enhanced cardiac muscle tissue network. |
| bio-central-nervous-system-top.mat | Enhanced central nervous system tissue network. |
| bio-chondrocyte-top.mat | Enhanced chondrocyte tissue network. |
| bio-embryo-top.mat | Enhanced embryo tissue network. |
| bio-epidermis-top.mat | Enhanced epidermis tissue network. |
| bio-heart-top.mat | Enhanced heart tissue network. |
| bio-lymphocyte-top.mat | Enhanced lymphocyte tissue network. |
| bio-muscle-top.mat | Enhanced muscle tissue network. |
| bio-natural-killer-cell-top.mat | Enhanced natural killer cell tissue network. |
| bio-nervous-system-top.mat | Enhanced nervous system tissue network. |
| bio-neuron-top.mat | Enhanced neuron tissue network. |
| bio-retina-top.mat | Enhanced retina tissue network |
| bio-skeletal-muscle-top.mat | Enhanced skeletal muscle tissue network. |
| bio-smooth-muscle-top.mat | Enhanced smooth muscle tissue network. |
| bio-t-lymphocyte-top.mat | Enhanced T lymphocyte tissue network. |
Study 2: Hi-C interaction networks
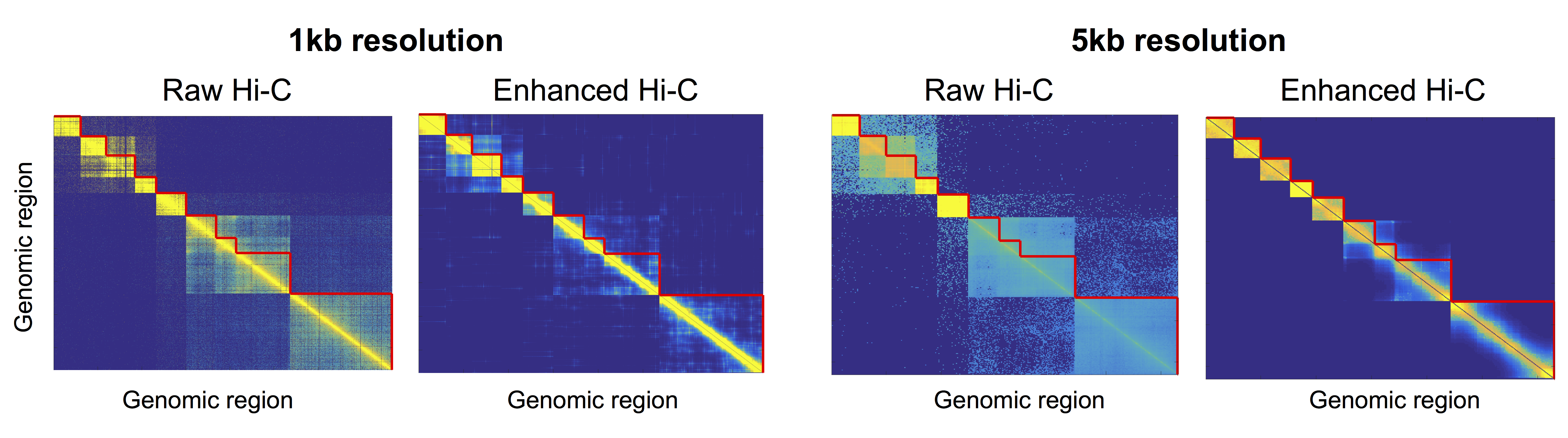
We applied NE to a Hi-C interaction networks. Hi-C is a 3C-based technology that allows measurement of pairwise chromatin interaction frequencies within a cell population. Hi-C read data can be thought of as a network where genomic regions are nodes and the normalized read counts mapped to two bins are weighted edges.
Visual inspection of the Hi-C contact matrix before and after Hi-C network is denoised using NE reveals an enhancement of edges within each community and sharper boundaries between communities (figure below). This improvement is particularly clear for the 5kb resolution data, where communities that were visually undetectable in the raw data become clear after denoising with NE.
Datasets
| File | Description |
|---|---|
| bio-1kb-preprocessed.zip | 1Kb resolution HiC data. The network was generated by stitching together sqrtVC normalized domains as described in the manuscript. The data was generated by Rao et al., Cell 2014. |
| bio-5kb-preprocessed.zip | 5Kb resolution HiC data. The network was generated by stitching together sqrtVC normalized domains as described in the manuscript. The data was generated by Rao et al., Cell 2014. |
| bio-1kb-enhanced.zip | 1Kb resolution Matlab .mat files containing the raw and NE-enhanced networks. NMI_all_1kresults.mat includes the NMI values for clustering raw and denoised networks. |
| bio-5kb-enhanced.zip | 5Kb resolution Matlab .mat files containing the raw and NE-enhanced networks. NMI_all_1kresults.mat includes the NMI values for clustering raw and denoised networks. |
Study 3: Butterfly species similarity networks
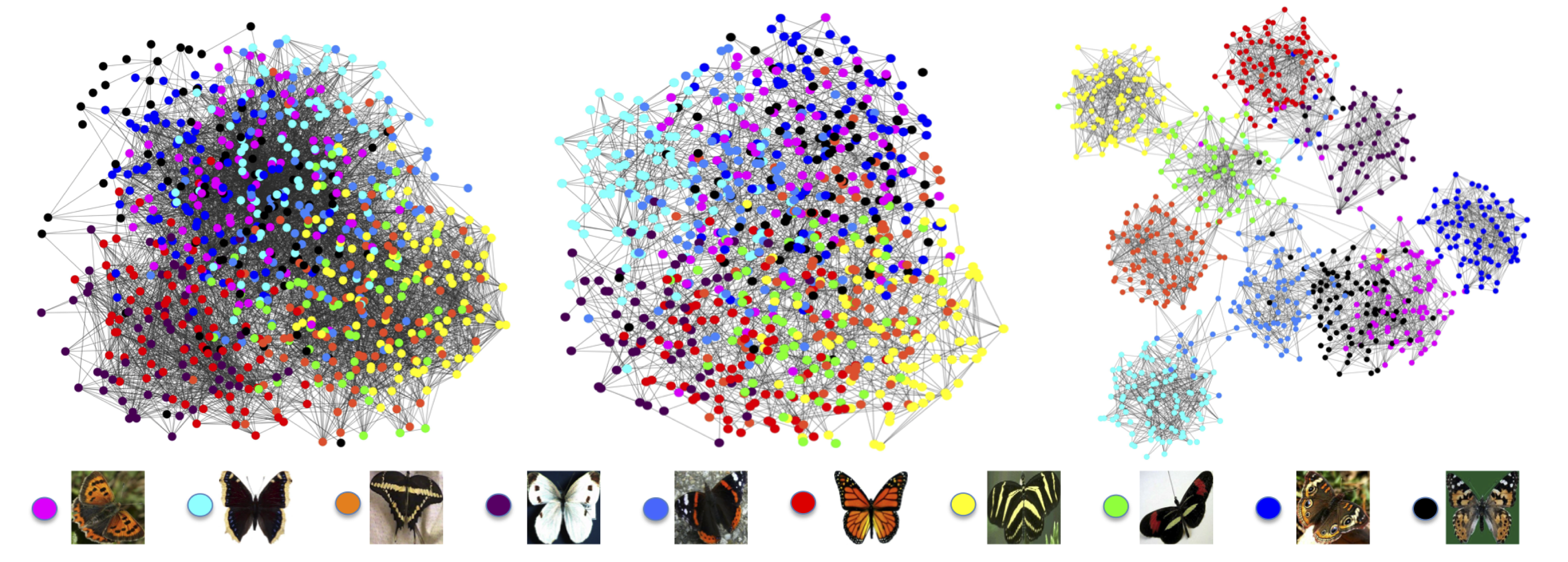
We applied NE to the Leeds butterfly fine-grained species image dataset. Fine-grained image retrieval aims to distinguish categories with subtle differences (e.g., monarch butterfly vs. peacock butterfly). We analyzed weighted similarity networks representing pairwise affinity between images of butterfly species.
Visual inspection indicates that NE is able to greatly improve the overall similarity network for fine-grain identification. Prior to NE, all the images are tangled together without a clear clustering (figure below, left). The resulting similarity network after applying NE clearly shows clusters representing different butterfly species (figure below, right).
Datasets
| File | Description |
|---|---|
| bio-raw-butterfly-network.mat.tar.gz | Weighted network representing image similarity of butterfly species. |
Network Enhancement code
You can download the code directly, which comes with sample biological network data. This is a Matlab implementation.